|
Kenneth T. He
|
| HOME | PROJECTS |
|
|
|
Born in Durham, NC I was born at Duke Children's Hospital & Health Center in Durham, NC. I have an elder sister and both of us grew up in Chapel Hill, NC. |

|

|
School I am attending Wake Forest University. I am a member of the swim team and a core member for our team. |
|
1. Developing a Tool for Reverse Complement DNA Sequences After having learned Java programming language in the past semester, I have developed a standalone and Web-based tool for reverse complement DNA sequences. For more details, please click here. |
|
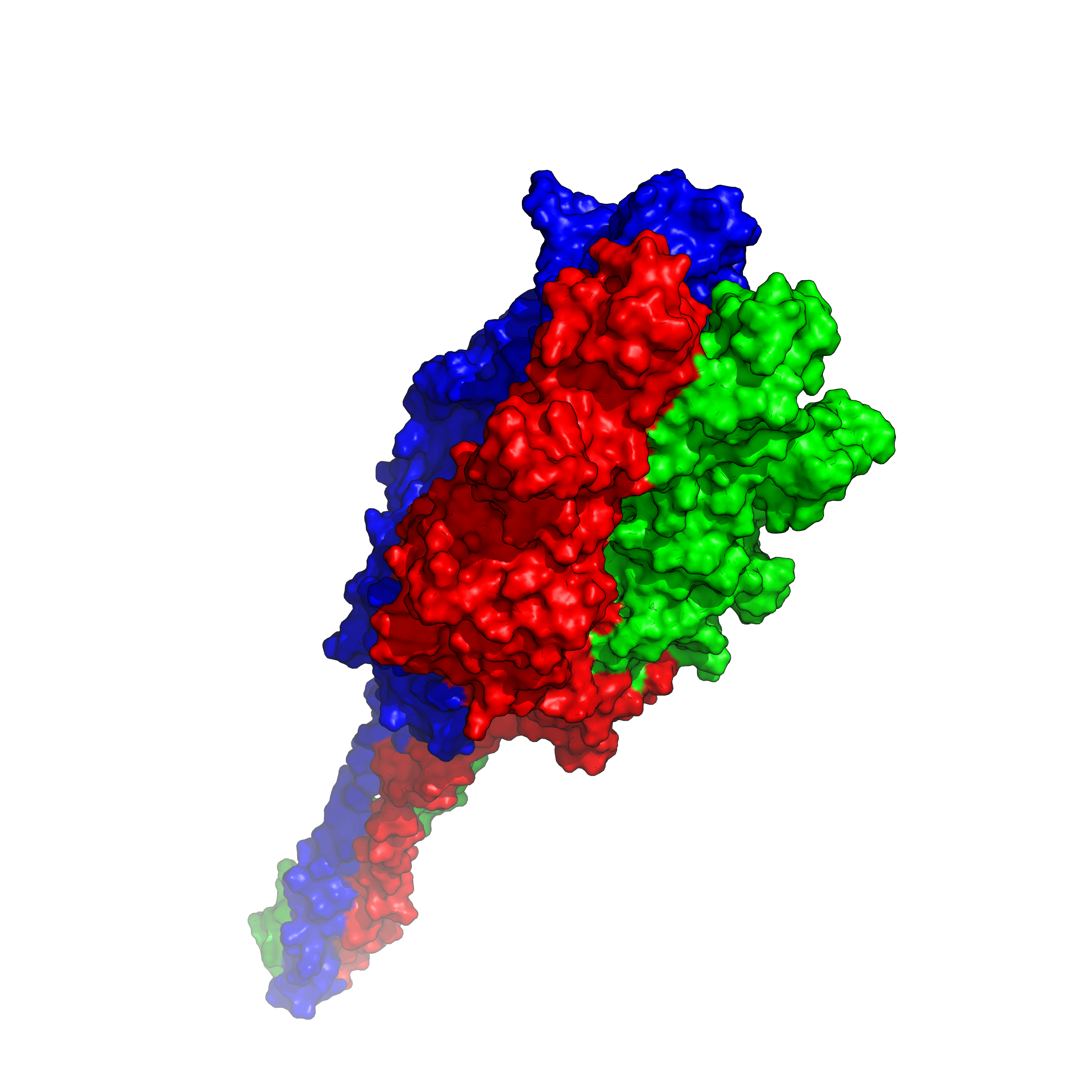
2. Predicting SARS-CoV-2 3D Structure Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a virus that causes respiratory illness in humans. It was first reported in December 2019 in an outbreak occurring in Wuhan, China. It is still spreading and affecting humans' working environment & life styles worldwide. Predicting its trimer 3D structure can be helpful in developing reliable vaccine and/or targeted therapies. |
|
|
| Since the spring of 2022, I got started to setup an AlphaFold2 running platform on my workstation with GPUs, on which I used to play games. With help from my dad, I have set AlphaFold2 up on my workstation and have run a number of tests for predicting protein structures based on their amino acid (AA) sequences. AlphaFold2 is an AI system developed by Google DeepMind that predicts a protein or complex's 3D structure from its AA sequence. It regularly achieves accuracy competitive with experiment. With the current ongoing virus of COVID-19/SARS-CoV-2, I have decided to work on this specific strand of protein and be able to estimate what it looks like. Using this application, I can visualize the SARS-CoV-2 structure and export different types of images including surface, sphere and cartoon. The protein of SARS-CoV-2 is a trimer. We can distinguish the three distinct strands through different colors to display these proteins using a program called PyMOL. |
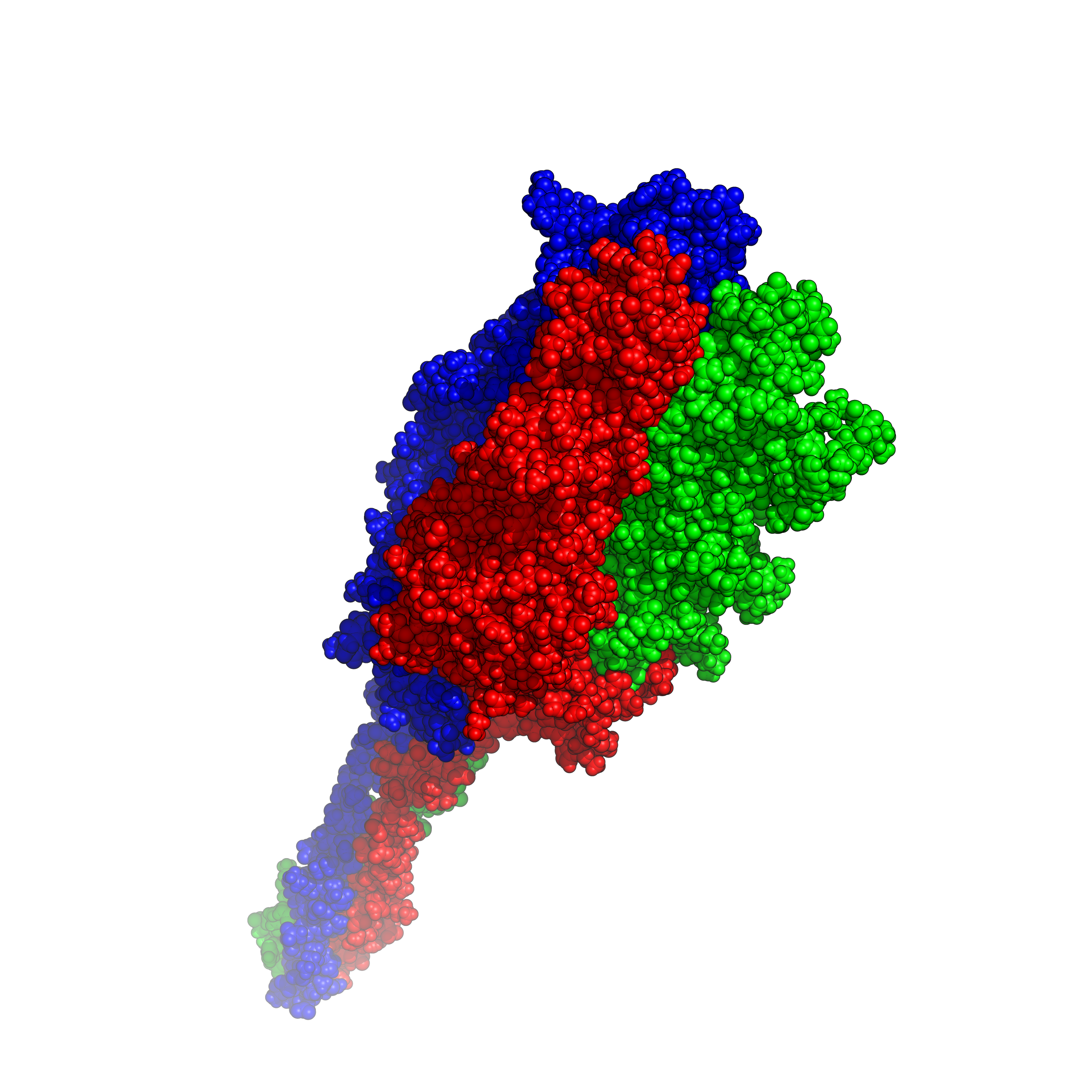
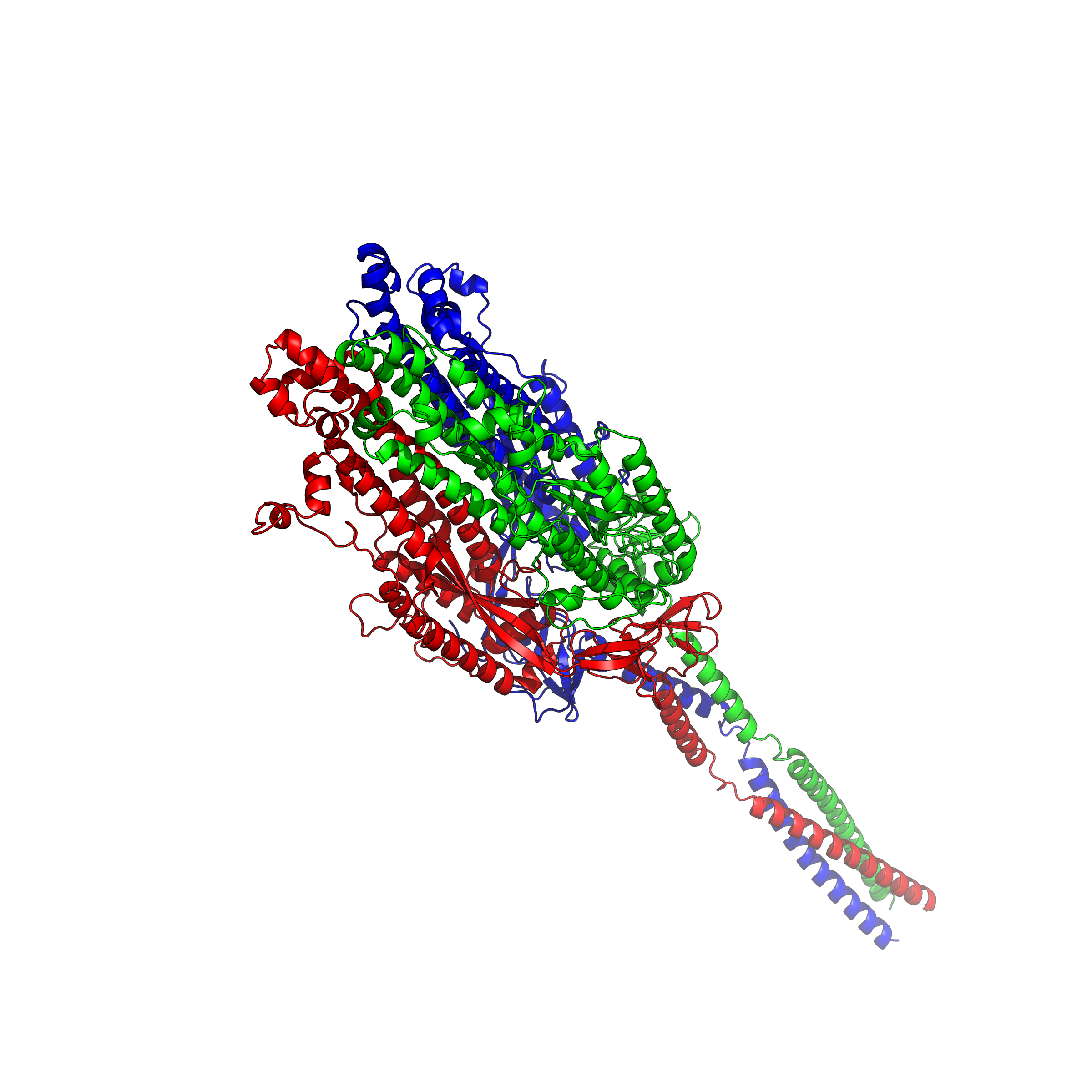
|